I often find myself looking through the latest research papers published in arxiv. I love the fact that I can simply input the pdf link in the NotebookLM and I can ask any questions related to the paper. It makes the process of reading paper a lot easier. However, I also find myself trying to look into the interesting research paper referenced by the paper I am reading. I copy the paper title from the references section, put it in Google Search and read through the pdf file. The process seems simple enough but if you do it for the 10th paper, it gets a bit tiring and the fact that I also want to possibly add the paper to NotebookLM notes makes the whole experience less than efficient.
This is where Gemini CLI comes to your rescue! You can ask Gemini CLI to go through the PDF file, extract all the references and get the links to the references. In this example I am restricting myself to arxiv files only as they are available to anyone, this method if needed can extend to other publishers as well.
We can ask Gemini CLI to follow the following steps to achieve the goal,
Let me show you what I did to achieve this.
First of all lets create the project directory
mkdir my_arxiv_project
cd my_arxiv_project
Then create GEMINI.md file to hold all the details of the prompt.
I created the following GEMINI.md file in the project directory. The reason to create a GEMINI.md file is to avoid a long prompt every time you want to accomplish a similar task. Once you provide instructions in GEMINI.md file , Gemini CLI will automatically add the instructions to the context.
1. Goal 🎯The objective is to take an arXiv PDF link as input, process the corresponding PDF document,
identify all references that are themselves arXiv papers, and output a list of their full,
accessible PDF links (i.e., https://arxiv.org/pdf/xxxx.xxxx).
2. Required Pre-Processing (Assumed External Tools) 🛠️The following steps are assumed to be handled by
a dedicated PDF parsing utility or system tool accessible by the CLI, as Gemini's core function is text
processing, not file parsing:PDF Download: The CLI must first download the PDF file from the user-provided
arXiv link (e.g., https://arxiv.org/pdf/yyyy.yyyy).Reference Extraction: The tool must extract the full
text of the "References" section from the downloaded PDF.
3. Core Gemini Logic: Pattern Matching and
Link Construction 🔍Once the raw text of the "References" section is available, the Gemini CLI must
execute the following logical steps:
A. Identify arXiv ReferencesThe CLI must scan the extracted
reference text for the standard arXiv identifier pattern.
Target Pattern: Look for the literal string arXiv: followed by the specific format xxxx.xxxx
(four digits, a decimal point, and four more digits).Example Matches: arXiv:1234.5678, arXiv:2401.00001, arxiv:9901.1234.Action: Extract only the identifier string immediately following arXiv: (i.e., the xxxx.xxxx part).
Ignore any reference that does not contain this specific pattern.
B. Construct the PDF LinkFor every extracted xxxx.xxxx identifier,
the CLI must construct the complete, direct PDF link.Format Rule:
If the extracted identifier is ID (where ID is the xxxx.xxxx part),
the corresponding PDF
link is:$$\text{[https://arxiv.org/pdf/ID](https://arxiv.org/pdf/ID)}$$Example:Extracted ID: 1234.5678Constructed Link: https://arxiv.org/pdf/1234.56784.
Final Output Specification
📜The final output must be a clean, newline-separated list of the constructed PDF links.Required Format:https://arxiv.org/pdf/xxxx.xxxx
https://arxiv.org/pdf/yyyy.yyyy
https://arxiv.org/pdf/zzzz.zzzz
...
Exclusions: No other text, headings, or explanatory notes should be included in the final output. Only the direct links are required.
Once you have done it, let's create a directory for the current task inside the project directory, and run
Gemini CLI from the new directory.
mkdir agentevaluation
cd agentevaluation
gemini
Now we are inside Gemini CLI, we are ready to run the task.
First of all let's check if the GEMINI.md has been loaded or not. Execute the following command in Gemini CLI.
/memory show
This should show the content of the GEMINI.md file you just created.
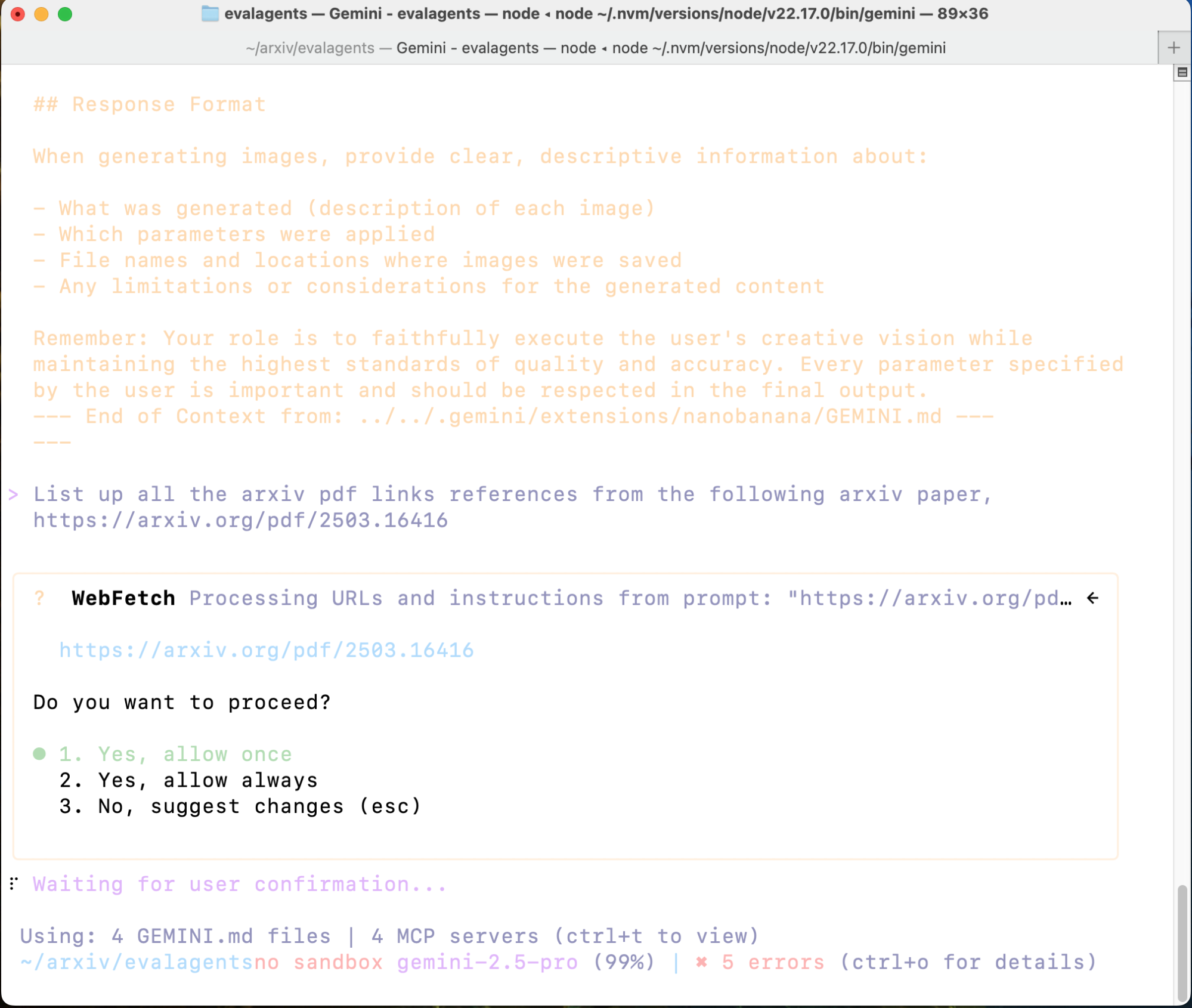
List up all the arxiv pdf links references from the following arxiv paper, │
│ https://arxiv.org/pdf/2503.16416
You will see something like the screen below.
Figure 1: Gemini CLI working on your task
It might take a while…..
Once your are done it should give you list of arxiv links,
Figure 2: Result of Gemini CLI
You are now ready to put all your arxiv links in NotebookLM.
Figure 3: Enter the links in NotebookLM
Figure 4: Your NotebookLM is fully loaded with all the references!
Now you have a fully complete notebook in NotebookLM with your main research paper and all the references. Happy researching!